Researchers introduce the “Telomouse”. By making a subtle genetic alteration in standard lab mice, they’ve made their telomeres, which protect the chromosome ends, more closely resemble those in humans. Telomeres are critical for maintaining genetic integrity and promoting healthy aging while reducing cancer risk. Standard lab mice have telomeres five times longer than humans, creating challenges in modeling their role in human aging and cancer. The Telomouse model, developed by incorporating a genetic variation from a mouse species with naturally shorter telomeres, provides a valuable resource for in-depth aging and cancer research, highlighting the importance of the RTEL1 protein in determining telomere length. This discovery promises to reveal new insights into the genetics of aging and may contribute to enhanced longevity and well-being.

[Jerusalem, Israel] – In an exciting scientific breakthrough, a team of researchers led by Professor Yehuda Tzfati from the Institute of Life Science at the Hebrew University and Professor Klaus Kaestner from the University of Pennsylvania Perelman School of Medicine, has introduced the “Telomouse.” This discovery involves changing just one tiny building block in one gene of ordinary lab mice, Mus musculus, to make their telomeres (our chromosome caps) look much more like the telomeres in humans.
Telomeres hold a pivotal responsibility in safeguarding our genetic material and ensuring the orderly division of our cells. Maintaining their structural integrity and optimal length holds the potential to diminish the risk of cancer and facilitate a healthier aging process. However, a significant hurdle has emerged: conventional laboratory mice possess telomeres approximately five times longer than those in humans. This disparity has posed a formidable challenge in using mice models for comprehending the implications of telomeres for human aging and cancer.
In the development of the Telomouse model, researchers turned their attention to a distinct mouse species, M. spretus, notable for its inherently shorter telomeres. Within the genetic code of these mice, a subtle variation within a pivotal protein known as RTEL1 was identified. By transferring this genetic distinction into typical laboratory mice, they succeeded in producing a lineage of mice with human-length telomeres. These novel Telomice exhibit robust health and reproductive capabilities, making them an exceptional resource for in-depth investigations into the complex realms of aging and cancer.
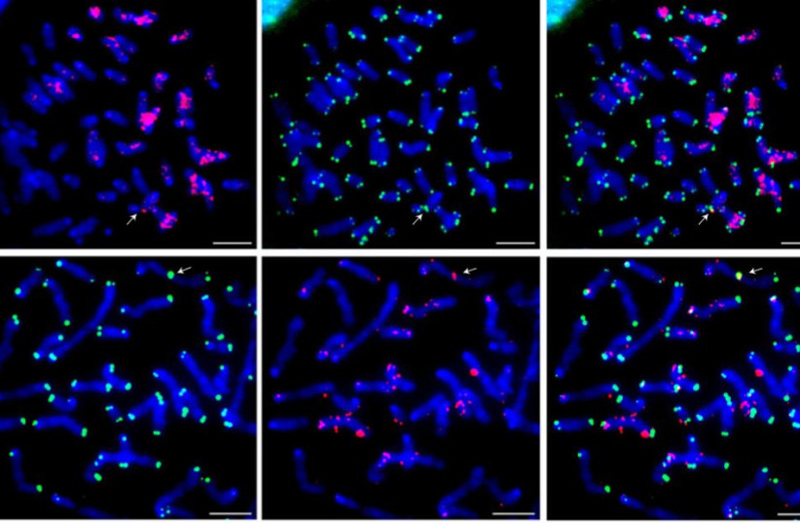
This study illuminates the central role of RTEL1 as the arbiter of telomere length. A nuanced modification to this crucial protein has enabled scientists to fashion a mouse model that closely approximates the human telomere length.
During the research, the researchers also achieved an invaluable breakthrough in our ability to measure the length of each single telomere, and particularly the shortest telomeres in the cell which are the ones to dictate the cellular function and fate. They developed a novel method for measuring the precise length of individual telomeres using a new generation of DNA sequencing called nanopore sequencing. This method, termed ‘NanoTelSeq’, enables evaluating the ‘telomeric health’ in samples from blood or other tissues of healthy individuals, as well as patients of cancers and aging diseases, and improve diagnosis, prognosis and treatment of these patients.
Professor Yehuda Tzfati, the principal investigator of this endeavor, posits, “The Telomouse model is promising to enrich our comprehension of the intricate nexus between telomeres, cancer and the aging process. I believe that NanoTelSeq will replace currently used methods, enable accurate evaluation of the telomere state in patients and healthy individuals, and reveal how it affects human health. Such insights will hopefully culminate in innovative strategies for combating cancer and fostering the well-being of aging individuals.”
The research paper titled “Telomouse — a mouse model with human-length telomeres generated by a single amino acid change in RTEL1” is now available in Nature Communications and can be read at https://t.ly/BsJlO.